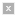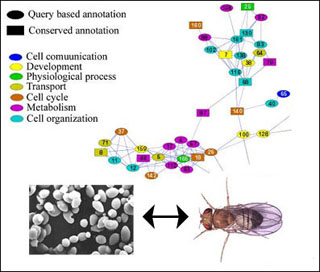
Techniques to model and analyze biological pathways are among the topics covered in this class. For instance, this map excerpt shows some protein interaction pathways in yeast S. cerevisiae that are also found in the fruit fly D. melanogaster: see assigned readings for the class. (Protein map excerpt courtesy of Tomer Shlomi et al. Source: Shlomi, T., et al. "QPath: A Method for Querying Pathways in a Protein-protein Interaction Network." BMC Bioinformatics 7 (2006): 199. Yeast and fly images courtesy of NASA.)
Instructor(s)
Prof. C. Forbes Dewey, Jr.
Prof. Hanry Yu
Prof. Sourav Saha Bhowmick
MIT Course Number
20.453J / 2.771J / HST.958J
As Taught In
Fall 2008
Level
Graduate
Course Description
Course Features
- Selected lecture notes
- Assignments: presentations with examples
- Assignments: programming (no examples)
- Assignments: written with examples
Course Description
This course teaches the design of contemporary information systems for biological and medical data. Examples are chosen from biology and medicine to illustrate complete life cycle information systems, beginning with data acquisition, following to data storage and finally to retrieval and analysis. Design of appropriate databases, client-server strategies, data interchange protocols, and computational modeling architectures. Students are expected to have some familiarity with scientific application software and a basic understanding of at least one contemporary programming language (e.g. C, C++, Java, Lisp, Perl, Python). A major term project is required of all students. This subject is open to motivated seniors having a strong interest in biomedical engineering and information system design with the ability to carry out a significant independent project.
This course was offered as part of the Singapore-MIT Alliance (SMA) program as course number SMA 5304.
Other Versions
Other OCW Versions
Archived versions: ![]()


